About the Gosling Project
The combination of diverse data types and analysis tasks in genomics has resulted in the development of a wide range of visualization techniques and tools. However, most existing tools are tailored to a specific problem or data type and offer limited customization, making it challenging to optimize visualizations for new analysis tasks or datasets. To address this challenge, we designed Gosling—a grammar for interactive and scalable genomics data visualization. Gosling balances expressiveness for comprehensive multi-scale genomics data visualizations with accessibility for domain scientists. Our accompanying JavaScript toolkit called Gosling.js provides scalable and interactive rendering. Gosling.js is built on top of an existing platform for web-based genomics data visualization to further simplify the visualization of common genomics data formats.
Why We Name This Project Gosling
"Gosling" has the following multiple meanings:
-
Gosling is the abbreviation of Grammar Of Scalable Linked Interactive Nucleotide Graphics.
-
Gosling also means baby goose 🐥. You can see a lot of baby geese during the spring in Boston, where our team is based.

-
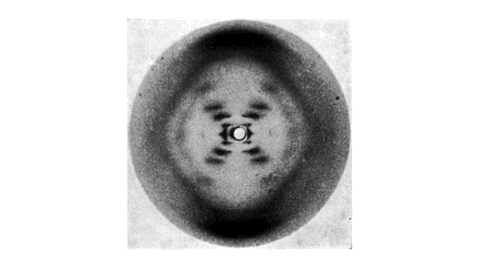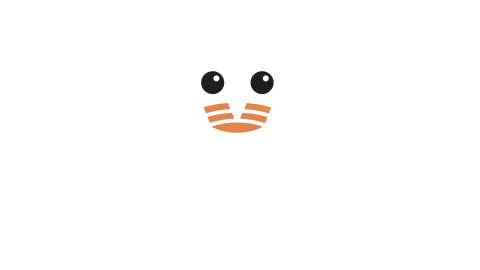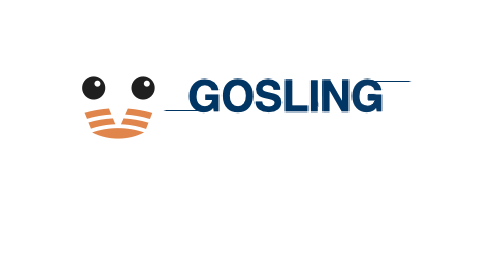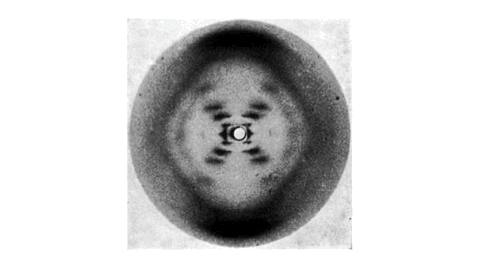
Most importantly, Gosling is named after a British scientist, Raymond Gosling, who took the X-ray diffraction image of DNA called Photo 51 under the supervision of Rosalind Franklin. This image visualized the structure of DNA and was taken as critical evidence in inferring the double-helix model of DNA. For more details about their achievement, read this long article.

Gosling Logo: A Baby Goose in the Photo 51
The logo of Gosling reflects both Photo 51 and a baby goose. Can you see the face of a baby goose in Photo 51?
How to Cite Gosling
@article{lyi2021gosling,
title={Gosling: A Grammar-based Toolkit for Scalable and Interactive Genomics Data Visualization},
author={Sehi L'Yi and Qianwen Wang and Fritz Lekschas and Nils Gehlenborg},
year={2021},
publisher={OSF Preprints},
url={osf.io/6evmb},
doi={10.31219/osf.io/6evmb},
}